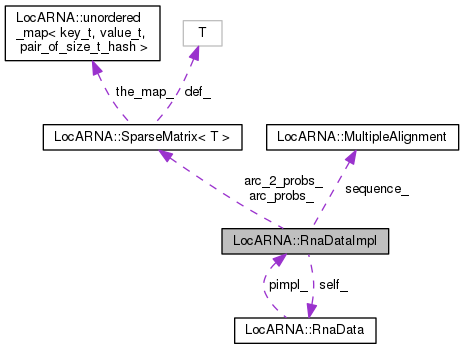
Implementation of RnaData.
More...
#include <rna_data_impl.hh>
|
| | RnaDataImpl (RnaData *self, const RnaData &rna_dataA, const RnaData &rna_dataB, const Alignment::edges_t &alignment, double p_expA, double p_expB) |
| | Construct as consensus of two aligned RNAs. More...
|
| |
| | RnaDataImpl (RnaData *self, double p_bpcut) |
| | Almost empty constructor. More...
|
| |
| void | init_from_fixed_structure (const SequenceAnnotation &structure, bool stacking) |
| | initialize from fixed structure More...
|
| |
| void | init_from_rna_ensemble (const RnaEnsemble &rna_ensemble, const PFoldParams &pfoldparams) |
| | initialize from rna ensemble More...
|
| |
| std::istream & | read_pp_sequence (std::istream &in) |
| | read sequence section of pp-format More...
|
| |
| std::istream & | read_pp_arc_probabilities (std::istream &in) |
| | read section of base pair probabilities of pp-format More...
|
| |
| std::ostream & | write_pp_sequence (std::ostream &out) const |
| | write section of base pair probabilities of pp-format More...
|
| |
| std::ostream & | write_pp_arc_probabilities (std::ostream &out, double p_outbpcut, bool stacking) const |
| | write section of base pair probabilities of pp-format More...
|
| |
| void | init_as_consensus_dot_plot (const Alignment::edges_t &edges, const RnaData &rna_dataA, const RnaData &rna_dataB, double p_expA, double p_expB, bool stacking) |
| | Initialize as consensus of two aligned RNAs. More...
|
| |
| double | consensus_probability (double pA, double pB, size_t sizeA, size_t sizeB, double p_expA, double p_expB) const |
| | Consensus probability. More...
|
| |
| void | drop_worst_bps (size_t keep) |
| | Drop base pairs with lowest probability. More...
|
| |
Implementation of RnaData.
Construct as consensus of two aligned RNAs.
- Parameters
-
| self | pointer to corresponding RnaData object |
| rna_dataA | data of RNA A |
| rna_dataB | data of RNA B |
| alignment | pairwise alignment of A and B |
| p_expA | background probability A |
| p_expB | background probability B |
| LocARNA::RnaDataImpl::RnaDataImpl |
( |
RnaData * |
self, |
|
|
double |
p_bpcut |
|
) |
| |
Almost empty constructor.
- Parameters
-
| self | pointer to corresponding RnaData object |
| p_bpcut | cutoff probability |
| double LocARNA::RnaDataImpl::consensus_probability |
( |
double |
pA, |
|
|
double |
pB, |
|
|
size_t |
sizeA, |
|
|
size_t |
sizeB, |
|
|
double |
p_expA, |
|
|
double |
p_expB |
|
) |
| const |
Consensus probability.
- Parameters
-
| pA | probability A |
| pB | probability B |
| sizeA | number of rows in sequence A |
| sizeB | number of rows in sequence B |
| p_expA | background probability A |
| p_expB | background probability B |
- Precondition
- p_bpcut_ is initialized
Essentially computes a weighted geometric mean; some care is taken, to avoid total extinction and to restrict the accumulation of (small) probabilities.
- Returns
- consensus probability
| void LocARNA::RnaDataImpl::drop_worst_bps |
( |
size_t |
keep | ) |
|
Drop base pairs with lowest probability.
- Parameters
-
| keep | the maximum number of base pairs to keep |
| void LocARNA::RnaDataImpl::init_as_consensus_dot_plot |
( |
const Alignment::edges_t & |
edges, |
|
|
const RnaData & |
rna_dataA, |
|
|
const RnaData & |
rna_dataB, |
|
|
double |
p_expA, |
|
|
double |
p_expB, |
|
|
bool |
stacking |
|
) |
| |
Initialize as consensus of two aligned RNAs.
- Parameters
-
| edges | alignment edges |
| rna_dataA | rna data A |
| rna_dataB | rna data B |
| p_expA | background probability A |
| p_expB | background probability B |
| stacking | if true, stacking consensus is computed |
| void LocARNA::RnaDataImpl::init_from_fixed_structure |
( |
const SequenceAnnotation & |
structure, |
|
|
bool |
stacking |
|
) |
| |
initialize from fixed structure
- Parameters
-
| structure | fixed structure |
| stacking | whether to initialize stacking terms |
| void LocARNA::RnaDataImpl::init_from_rna_ensemble |
( |
const RnaEnsemble & |
rna_ensemble, |
|
|
const PFoldParams & |
pfoldparams |
|
) |
| |
initialize from rna ensemble
- Parameters
-
| rna_ensemble | rna ensemble |
| pfoldparams | folding parameters. if stacking, initialize stacking terms; if noLP, drop lonely pairs |
| std::istream & LocARNA::RnaDataImpl::read_pp_arc_probabilities |
( |
std::istream & |
in | ) |
|
read section of base pair probabilities of pp-format
- Parameters
-
Reads only base pairs with probabilities greater than p_bpcut_; reads stacking only if has_stacking_
| std::istream & LocARNA::RnaDataImpl::read_pp_sequence |
( |
std::istream & |
in | ) |
|
read sequence section of pp-format
- Parameters
-
- Returns
- stream
this section comprises sequence/multiple alignment (possibly including sequence anchor annotation)
| std::ostream & LocARNA::RnaDataImpl::write_pp_arc_probabilities |
( |
std::ostream & |
out, |
|
|
double |
p_outbpcut, |
|
|
bool |
stacking |
|
) |
| const |
write section of base pair probabilities of pp-format
Write arc probabilities.
- Parameters
-
| out | ouput stream |
| p_outbpcut | cutoff probabilitiy |
| stacking | whether to write stacking probabilities; if stacking but !has_stacking_, no stacking terms are written but flag #STACKS is written to output |
- Returns
- stream
Write only base pairs with probabilities greater than p_outbpcut
Writes arc and stacking probabilities to stream; filters by probability threshold p_outbpcut
| std::ostream & LocARNA::RnaDataImpl::write_pp_sequence |
( |
std::ostream & |
out | ) |
const |
write section of base pair probabilities of pp-format
- Parameters
-
- Returns
- stream
sparse array for all probabilities that a pair (i,j) and its immediately inner pair (i+1,j-1) are formed simultaneously above threshold; analogous to arc_probs_
- Note
- arc_2_probs_ has entry (i,j) implies arc_probs_ has entry (i,j)
sparse array for all arc probabilities above threshold; the array is used when reading in the probabilities and for merging probs during pp-output
| RnaData* LocARNA::RnaDataImpl::self_ |
- pointer to corresponding non-impl object
The documentation for this class was generated from the following files:
 1.8.11
1.8.11