|
LocARNA-1.8.11
|
|
LocARNA-1.8.11
|
class ArcMatches with additional mapping More...
#include <arc_matches.hh>
Public Member Functions | |
| ArcMatchesIndexed (const Sequence &seqA_, const Sequence &seqB_, const std::string &arcmatch_scores_file, int probability_scale, size_type max_length_diff, size_type max_diff_at_am, const MatchController &trace_controller, const AnchorConstraints &constraints) | |
| construct with explicit arc match score list More... | |
| ArcMatchesIndexed (const RnaData &rnadataA, const RnaData &rnadataB, double min_prob, size_type max_length_diff, size_type max_diff_at_am, const MatchController &trace_controller, const AnchorConstraints &constraints) | |
| construct from single base pair probabilities. More... | |
| const ArcMatch::idx_type | invalid_am_index () const |
| the invalid arc match index More... | |
| const ArcMatch::idx_type | am_index (const size_type &arcAIdx, const size_type &arcBIdx) const |
| Lookup arc match index by pair of arc indices. More... | |
| const ArcMatch & | am_index (const Arc &arcA, const Arc &arcB) const |
| Lookup arc match by pair of arcs. More... | |
 Public Member Functions inherited from LocARNA::ArcMatches Public Member Functions inherited from LocARNA::ArcMatches | |
| ArcMatches (const Sequence &seqA_, const Sequence &seqB_, const std::string &arcmatch_scores_file, int probability_scale, size_type max_length_diff, size_type max_diff_at_am, const MatchController &trace_controller, const AnchorConstraints &constraints) | |
| construct with explicit arc match score list More... | |
| ArcMatches (const RnaData &rnadataA, const RnaData &rnadataB, double min_prob, size_type max_length_diff, size_type max_diff_at_am, const MatchController &trace_controller, const AnchorConstraints &constraints) | |
| construct from single base pair probabilities. More... | |
| ~ArcMatches () | |
| clean up base pair objects | |
| void | read_arcmatch_scores (const std::string &arcmatch_scores_file, int probability_scale) |
| Reads scores for arc matches. More... | |
| void | write_arcmatch_scores (const std::string &arcmatch_scores_file, const Scoring &scoring) const |
| const BasePairs & | get_base_pairsA () const |
| returns the base pairs object for RNA A | |
| const BasePairs & | get_base_pairsB () const |
| returns the base pairs object for RNA B | |
| bool | explicit_scores () const |
| true, if arc match scores are explicit (because they are read in from a list) | |
| void | make_scores_explicit (const Scoring &scoring) |
| Make arcmatch scores explicit. More... | |
| score_t | get_score (const ArcMatch &am) const |
| size_type | num_arc_matches () const |
| total number of arc matches | |
| const ArcMatch & | arcmatch (size_type idx) const |
| get arc match by its index | |
| const ArcMatchIdxVec & | common_right_end_list (size_type i, size_type j) const |
| list of all arc matches that share the common right end (i,j) | |
| const ArcMatchIdxVec & | common_left_end_list (size_type i, size_type j) const |
| list of all arc matches that share the common left end (i,j) | |
| void | get_max_right_ends (size_type al, size_type bl, size_type *max_ar, size_type *max_br, bool no_lonely_pairs) const |
| get the maximal right ends of any arc match with left ends (al,bl). More... | |
| void | get_min_right_ends (size_type al, size_type bl, size_type *min_ar, size_type *min_br) const |
| bool | exists_inner_arc_match (const ArcMatch &am) const |
| const ArcMatch & | inner_arc_match (const ArcMatch &am) const |
| void | sort_right_adjacency_lists () |
| const_iterator | begin () const |
| begin of arc matches vector | |
| const_iterator | end () const |
| end of arc matches vector | |
Additional Inherited Members | |
 Public Types inherited from LocARNA::ArcMatches Public Types inherited from LocARNA::ArcMatches | |
| typedef std::vector< int >::size_type | size_type |
| size | |
| typedef BasePairs__Arc | Arc |
| arc | |
| typedef ArcMatchVec::const_iterator | const_iterator |
| const iterator over arc matches | |
 Protected Member Functions inherited from LocARNA::ArcMatches Protected Member Functions inherited from LocARNA::ArcMatches | |
| bool | is_valid_arcmatch (const Arc &arcA, const Arc &arcB) const |
| void | init_inner_arc_matchs () |
| initialize the vector of inner arc match indices | |
 Protected Attributes inherited from LocARNA::ArcMatches Protected Attributes inherited from LocARNA::ArcMatches | |
| size_type | lenA |
| length of sequence A | |
| size_type | lenB |
| length of sequence B | |
| BasePairs * | bpsA |
| base pairs of RNA A | |
| BasePairs * | bpsB |
| base pairs of RNA B | |
| size_type | max_length_diff |
| for max-diff-am heuristics | |
| size_type | max_diff_at_am |
| for max diff at arc matches heuristics | |
| const MatchController & | match_controller |
| allowed alignment traces by max-diff heuristics | |
| const AnchorConstraints & | constraints |
| for constraints | |
| bool | maintain_explicit_scores |
| whether scores are maintained explicitely or computed from pair probabilities | |
| ArcMatchVec | arc_matches_vec |
| vector of all maintained arc matches | |
| size_type | number_of_arcmatches |
| std::vector< score_t > | scores |
| vector of scores (of arc matches with the same index) | |
| Matrix< ArcMatchIdxVec > | common_right_end_lists |
| for each (i,j) maintain vector of the indices of the arc matchs that share the common right end (i,j) | |
| Matrix< ArcMatchIdxVec > | common_left_end_lists |
| for each (i,j) maintain vector of the indices of the arc matchs that share the common left end (i,j) | |
| ArcMatchIdxVec | inner_arcmatch_idxs |
| vector of indices of inner arc matches | |
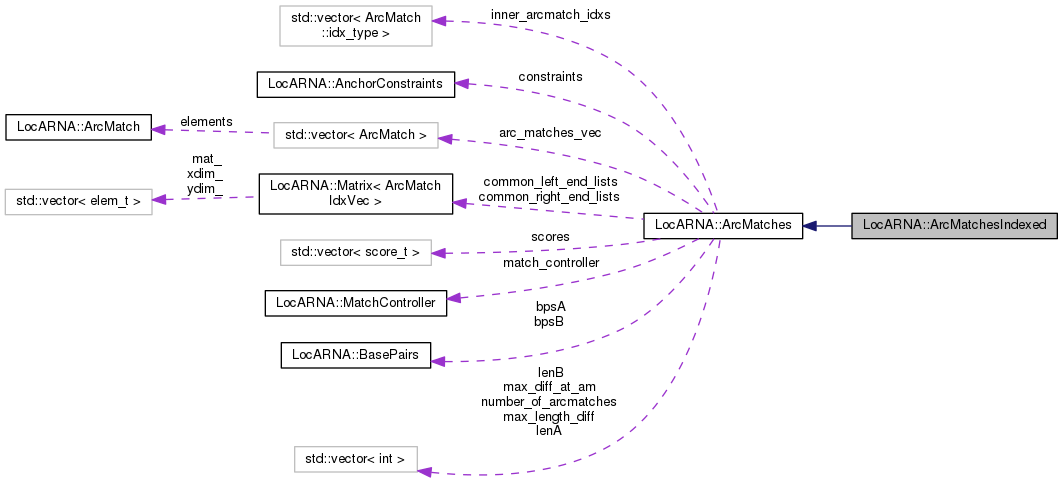
class ArcMatches with additional mapping
Like ArcMatches, maintain the relevant arc matches and their scores. Additionally, build an index to support mapping from pairs of arcs to arc matches.
|
inline |
construct with explicit arc match score list
construct from seqnames and explicit list of all scored arc matches together with their score.
| seqA_ | sequence A |
| seqB_ | sequence B |
| arcmatch_scores_file | file containing arc match scores |
| probability_scale | if >=0 read probabilities and multiply them by probability_scale |
| max_length_diff | accept arc matches only up to maximal length difference |
| trace_controller | accept only due to trace controller |
| constraints | accept only due to constraints |
|
inline |
construct from single base pair probabilities.
In this case, the object filters for relevant base pairs/arcs by min_prob. Registers constraints and heuristics and then calls read_arcmatch_scores. Constructs BasePairs objects for each single object and registers them. Generates adjacency lists of arc matches for internal use and sorts them. Lists contain only valid arc matches according to constraints and heuristics (see is_valid_arcmatch()). The constructed object does not explicitely represent/maintain the scores of arc matchs.
| rnadataA | data for RNA A |
| rnadataB | data for RNA B |
| min_prob | consider only arcs with this minimal probability |
| max_length_diff | consider arc matches only up to maximal length difference |
| trace_controller | arc matches only due to trace controller |
| constraints | arc matches only due to constraints |
|
inline |
Lookup arc match index by pair of arc indices.
| arcAIdx | index of arc in A |
| arcBIdx | index of arc in B |
|
inline |
Lookup arc match by pair of arcs.
| arcA | arc in A |
| arcB | arc in B |
|
inline |
the invalid arc match index
 1.8.11
1.8.11