Supplement to LocARNA-P: Accurate Boundary Prediction and Improved Detection of Structural RNAs
Sebastian Will, Tejal Joshi, Ivo L. Hofacker, Peter F. Stadler, Rolf Backofen
LocARNA-P Web-Server, Software Package, and Documentation
- Web server
The main functionality of LocARNA-P is made available via a web based interface by
the Freiburg RNA Tools. In order to activate the LocARNA-P functionality in the LocARNA server, please check the option "Probabilistic" in the advanced options. The option is available at the bottom of the input page after selecting "Show Advanced".
- Stand alone software and source code
LocARNA-P is implemented as part of the
LocARNA
software package
. LocARNA-P's multiple alignment functionality is
accessible via mlocarna --probabilistic. Furthermore,
the package contains scripts for refining genome-wide non-coding
RNA screens and the tool reliability-profile producing
the reliability profile plot and signal predictions.
- Documentation of LocARNA-P (as part of the LocARNA package) is available as PDF.
LocARNA-P Refinement of RNAz Boundaries
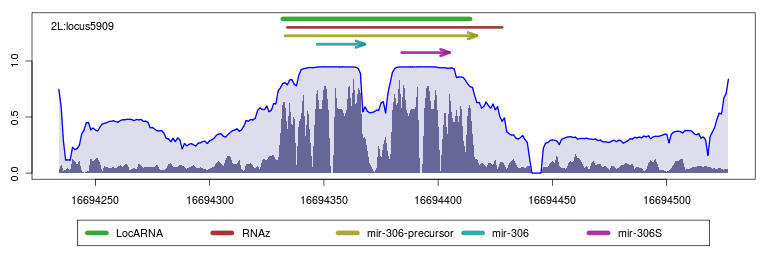
- Browse RNAz hits refined by LocARNA-P
Interactively view all reliability profiles of loci with LocARNA-P predictions, RNAz predictions and annotations.
- Raw data: Table of RNAz hits with annotation
Line format: locus chromosome start_RNAz end_RNAz orientation_RNAz id_annotation1 name_annotation1 start_annotation1 end_annotation1 orientation_annotation1 ...
- Raw data: Table of RNAz hits with LocARNA-P prediction and annotation
Line format: chromosome:locus start_RNAz end_RNAz orientation_RNAz start_LocARNA-P end_LocARNA-P orientation_LocARNA-P start_annotation end_annotation orientation_annotation on_value off_value
- orientation_LocARNA-P tells whether plus or minus strand was used for predicting the position.
- on_value and off_value are the repective values for fitting the two-step function.